Iterative Deepening DFS in R
Posted: , Last Updated:
bottom_reached = FALSE # Variable to keep track if we have reached the bottom of the tree
iterative_deepening_dfs <- function(start, target){
#' Implementation of iterative deepening DFS (depth-first search) algorithm to find the shortest path from a start to a target node..
#' Given a start node, this returns the node in the tree below the start node with the target value (or null if it doesn't exist)
#' Runs in O(n), where n is the number of nodes in the tree, or O(b^d), where b is the branching factor and d is the depth.
#' @param start the node to start the search from
#' @param target the value to search for
#' @return The node containing the target value or null if it doesn't exist.
# Start by doing DFS with a depth of 1, keep doubling depth until we reach the "bottom" of the tree or find the node we're searching for
depth = 1
while(!bottom_reached){
bottom_reached <<- TRUE # One of the "end nodes" of the search with this depth has to still have children and set this to false again
# One of the "end nodes" of the search with this depth has to still have children and set this to FALSE again
result = iterative_deepening_dfs_rec(start, target, 0, depth)
if(!is.null(result)){
# We've found the goal node while doing DFS with this max depth
return (result)
}
# We haven't found the goal node, but there are still deeper nodes to search through
depth = depth * 2
cat("Increasing depth to ", depth, "\n")
print(bottom_reached)
}
# Bottom reached is TRUE.
# We haven't found the node and there were no more nodes that still have children to explore at a higher depth.
return (NULL)
}
iterative_deepening_dfs_rec <- function(node, target, current_depth, max_depth) {
cat("Visiting Node ", node$value, "\n")
if(node$value == target){
# We have found the goal node we we're searching for
print("Found the node we're looking for!")
return (node)
}
if(current_depth == max_depth) {
print("Current maximum depth reached, returning...")
# We have reached the end for this depth...
if(length(node$children) > 0){
# ...but we have not yet reached the bottom of the tree
bottom_reached <<- FALSE
}
return (NULL)
}
# Recurse with all children
for (i in seq_along(node$children)){
result = iterative_deepening_dfs_rec(node$children[[i]], target, current_depth + 1, max_depth)
if(!is.null(result)){
# We've found the goal node while going down that child
return (result)
}
}
# We've gone through all children and not found the goal node
return (NULL)
}
About the algorithm and language used in this code snippet:
Iterative Deepening Depth-First Search Algorithm
The Iterative Deepening Depth-First Search (also ID-DFS) algorithm is an algorithm used to find a node in a tree. This means that given a tree data structure, the algorithm will return the first node in this tree that matches the specified condition. Nodes are sometimes referred to as vertices (plural of vertex) - here, we’ll call them nodes. The edges have to be unweighted. This algorithm can also work with unweighted graphs if mechanism to keep track of already visited nodes is added.
Description of the Algorithm
The basic principle of the algorithm is to start with a start node, and then look at the first child of this node. It then looks at the first child of that node (grandchild of the start node) and so on, until a node has no more children (we’ve reached a leaf node). It then goes up one level, and looks at the next child. If there are no more children, it goes up one more level, and so on, until it find more children or reaches the start node. If hasn’t found the goal node after returning from the last child of the start node, the goal node cannot be found, since by then all nodes have been traversed.
So far this has been describing Depth-First Search (DFS). Iterative deepening adds to this, that the algorithm not only returns one layer up the tree when the node has no more children to visit, but also when a previously specified maximum depth has been reached. Also, if we return to the start node, we increase the maximum depth and start the search all over, until we’ve visited all leaf nodes (bottom nodes) and increasing the maximum depth won’t lead to us visiting more nodes.
Specifically, these are the steps:
- For each child of the current node
- If it is the target node, return
- If the current maximum depth is reached, return
- Set the current node to this node and go back to 1.
- After having gone through all children, go to the next child of the parent (the next sibling)
- After having gone through all children of the start node, increase the maximum depth and go back to 1.
- If we have reached all leaf (bottom) nodes, the goal node doesn’t exist.
Example of the Algorithm
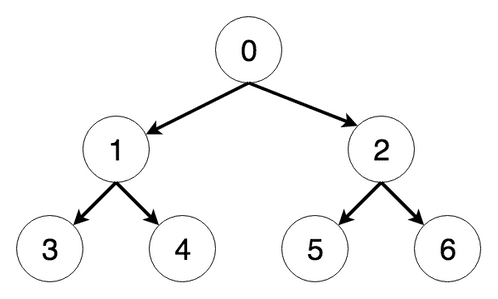
Consider the following tree:
The steps the algorithm performs on this tree if given node 0 as a starting point, in order, are:
- Visiting Node 0
- Visiting Node 1
- Current maximum depth reached, returning…
- Visiting Node 2
- Current maximum depth reached, returning…
- Increasing depth to 2
- Visiting Node 0
- Visiting Node 1
- Visiting Node 3
- Current maximum depth reached, returning…
- Visiting Node 4
- Current maximum depth reached, returning…
- Visiting Node 2
- Visiting Node 5
- Current maximum depth reached, returning…
- Visiting Node 6
- Found the node we’re looking for, returning…
Runtime of the Algorithm
If we double the maximum depth each time we need to go deeper, the runtime complexity of Iterative Deepening Depth-First Search (ID-DFS) is the same as regular Depth-First Search (DFS), since all previous depths added up will have the same runtime as the current depth (1/2 + 1/4 + 1/8 + … < 1). The runtime of regular Depth-First Search (DFS) is O(|N|) (|N| = number of Nodes in the tree), since every node is traversed at most once. The number of nodes is equal to b^d, where b is the branching factor and d is the depth, so the runtime can be rewritten as O(b^d).
Space of the Algorithm
The space complexity of Iterative Deepening Depth-First Search (ID-DFS) is the same as regular Depth-First Search (DFS), which is, if we exclude the tree itself, O(d), with d being the depth, which is also the size of the call stack at maximum depth. If we include the tree, the space complexity is the same as the runtime complexity, as each node needs to be saved.
R
R is an interpreted language first released in 1993 with a significant increase in popularity in recent years. It is primarily used for data mining and -science as well as statistics, and is a popular language in non-computer science disciplines ranging from Biology to Physics. R is dynamically typed, and has one of the widest variety of libraries for statistics, machine learning, data mining etc.
Getting to “Hello World” in R
The most important things first - here’s how you can run your first line of code in R.
- Download and install the latest version of R from r-project.org. You can also download an earlier version if your use case requires it.
- Open a terminal, make sure the
Rcommand is working, and that the command your’re going to be using is referring to the version you just installed by runningR --version. If you’re getting a “command not found” error (or similar), try restarting your command line, and, if that doesn’t help, your computer. If the issue persists, here are some helpful StackOverflow questions for Windows, Mac and Linux. - As soon as that’s working, you can run the following snippet:
print("Hello World"). You have two options to run this: 3.1 RunRin the command line, just paste the code snippet and press enter (PressCTRL + Dand typenfollowed by enter to exit). 3.2 Save the snippet to a file, name it something ending with.R, e.g.hello_world.R, and runRscript hello_world.R. Tip: use thelscommand (dirin Windows) to figure out which files are in the folder your command line is currently in.
That’s it! Notice how printing something to the console is just a single line in R - this low entry barrier and lack of required boilerplate code is a big part of the appeal of R.
Fundamentals in R
To understand algorithms and technologies implemented in R, one first needs to understand what basic programming concepts look like in this particular language.
Variables and Arithmetic
Variables in R are really simple, no need to declare a datatype or even declare that you’re defining a variable; R knows this implicitly. R is also able to easily define objects and their property, in multiple different ways.
some_value = 10
my_object <- list(my_value = 4)
attr(my_object, 'other_value') <- 3
print((some_value + my_object$my_value + attr(my_object, 'other_value'))) # Prints 17Arrays
Working with arrays is similarly simple in R:
# Create 2 vectors of length 3
vector1 <- c(1,2,3)
vector2 <- c(4,5,6)
# Create names for rows and columns (optional)
column.names <- c("column_1","column_2","column_3")
row.names <- c("row_1","row_2")
# Concatenate the vectors (as rows) to form an array, providing dimensions and row/column names
result <- array(c(vector1,vector2), dim = c(2,3), dimnames = list(row.names, column.names))
print(result)
# Prints:
# column_1 column_2 column_3
# row_1 1 3 5
# row_2 2 4 6As those of you familiar with other programming language like Java might have already noticed, those are not native arrays, but rather lists dressed like arrays. This means that arrays in R are considerably slower than in lower level programming languages. This is a trade off R makes in favor of simplicity. There are, however, packages which implement real arrays that are considerably faster.
Conditions
Just like most programming languages, R can do if-else statements:
value = 1
if(value==1){
print("Value is 1")
} else if(value==2){
print("Value is 2")
} else {
print("Value is something else")
}R can also do switch statements, although they are implemented as a function, unlike in other languages like Java:
x <- switch(
1,
"Value is 1",
"Value is 2",
"Value is 3"
)
print(x)Note that this function is pretty useless, but there are other functions for more complex use cases.
Loops
R supports both for and while loops as well as break and next statements (comparable to continue in other languages).
Additionally, R supports repeat-loops, which are comparable to while(true) loops in other languages, but simplify the code a little bit.
value <- 0
repeat {
value <- value + 1
if(value > 10) {
break
}
}
print(value)
value <- 0
while (value <= 10) {
value = value + 1
}
print(value)
value <- c("Hello","World","!")
for ( i in value) {
print(i)
}
for(i in 1:10){
print(i)
}Functions
Functions in R are easily defined and, for better or worse, do not require specifying return or arguments types. Optionally, a default for arguments can be specified:
my_func <- function (
a = "World"
) {
print(a)
return("!")
}
my_func("Hello")
print(my_func())(This will print “Hello”, “World”, and then ”!“)
Syntax
R requires the use of curly brackets ({}) to surround code blocks in conditions, loops, functions etc.;
While this can lead to some annoying syntax errors, it also means the use of whitespace for preferred formatting (e.g. indentation of code pieces) does not affect the code.
Advanced Knowledge of R
For more information, R has a great Wikipedia article. The official website is r-project.org.